Neural Optimism for Genetic Perturbation Experiments
Efficient data acquisition for genetic experiments using neural networks.
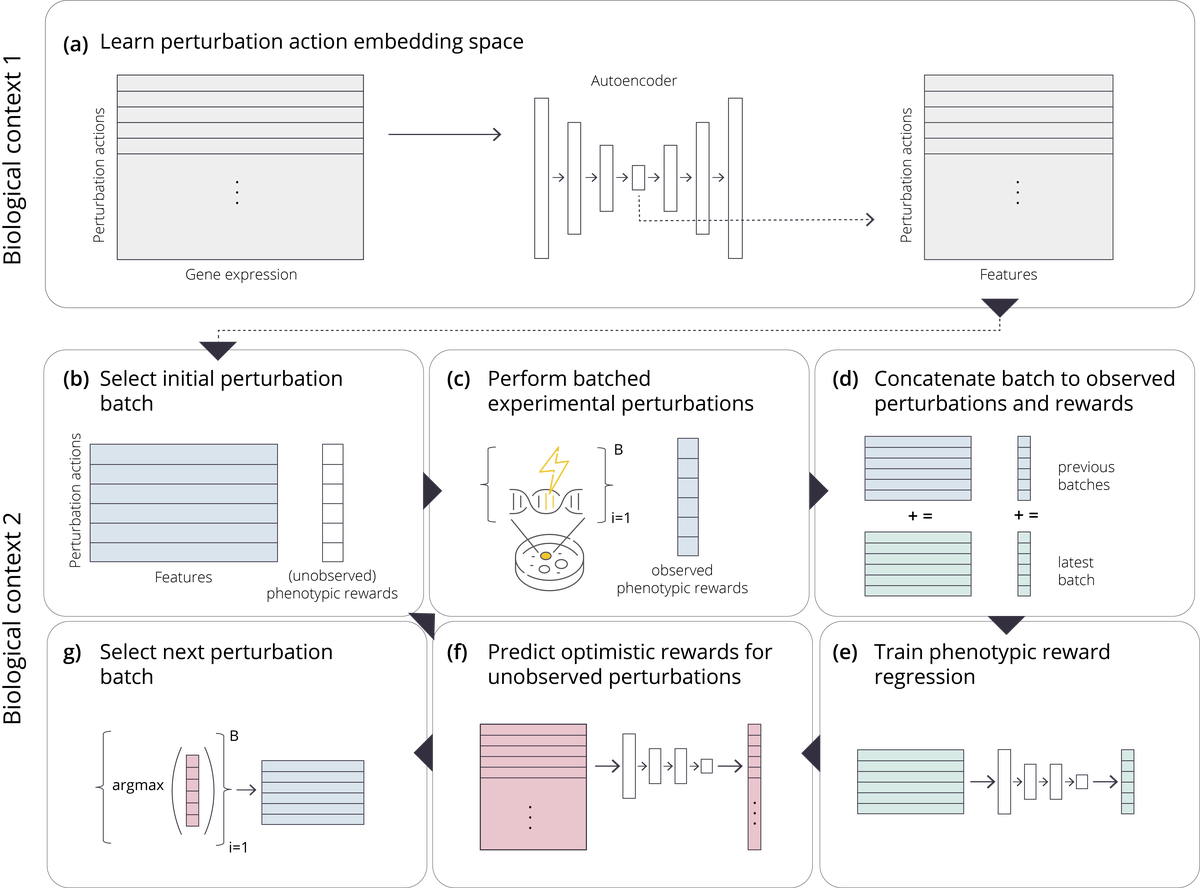
Overview
The problem of how to genetically modify cells in order to maximize a certain cellular phenotype has taken center stage in drug development over the last few years (with, for example, genetically edited CAR-T, CAR-NK, and CAR-NKT cells entering cancer clinical trials). Exhausting the search space for all possible genetic edits (perturbations) or combinations thereof is infeasible due to cost and experimental limitations. This work provides a theoretically sound framework for iteratively exploring the space of perturbations in pooled batches in order to maximize a target phenotype under an experimental budget. Inspired by this application domain, we study the problem of batch query bandit optimization and introduce the Optimistic Arm Elimination (OAE) principle designed to find an almost optimal arm under different functional relationships between the queries (arms) and the outputs (rewards). In this work we analyze several tractable mechanisms for optimistic gene perturbation discovery that use neural network function approximation. We analyze the convergence properties of OAE by relating it to the Eluder dimension of the algorithm’s function class and validate that OAE outperforms other strategies in finding optimal actions in experiments on simulated problems, public datasets well-studied in bandit contexts, and in genetic perturbation datasets when the regression model is a deep neural network. OAE also outperforms the benchmark algorithms in 3 of 4 datasets in the GeneDisco experimental planning challenge. Our algorithms can be used in problems beyond the setting of genetic perturbation experiments.